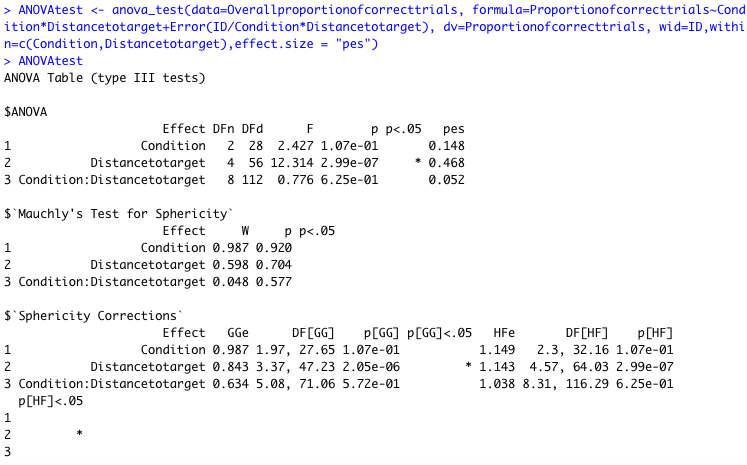
It's not ideal and you have to make it very very clear that those error bars are educated guesses but maybe is a fair way of using the data you have. After further reading I found the function Anova () in car package can be used to compute two-way ANOVA test for unbalanced designs.
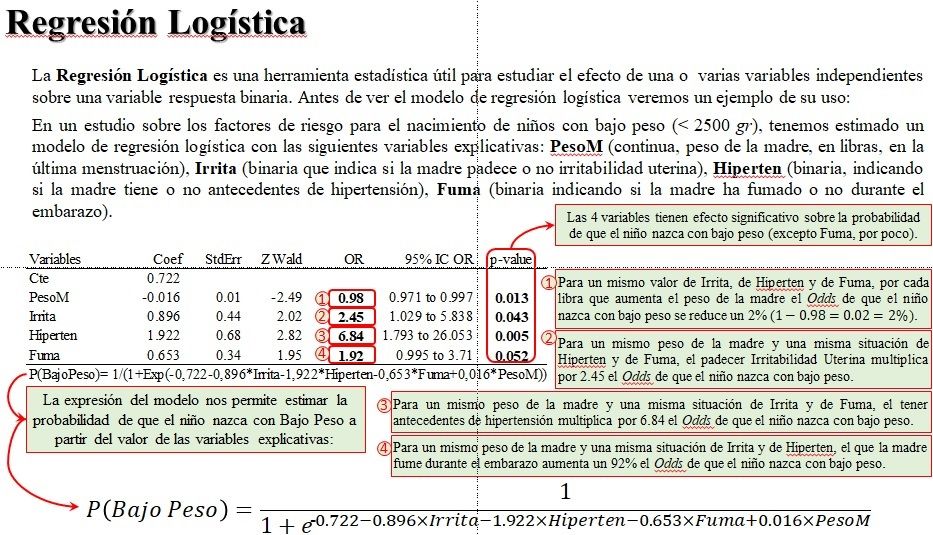
Whereas the difference between "no glu" and glu is sufficiently large to be worth discussing. This suggests that you cannot say much about the difference between "no glu" and gluFCCP and gluNV. Segments(x0= b, y0= dat$down, y1= dat$up) Internal_concentration = c(2.38, 8.57, 3.42, 6.17, 4.58,3.51))ĭat <- datĭat$up <- dat $internal_concentration + 1ĭat$down <- dat$internal_concentration - 1ī <- barplot(dat $internal_concentration, names.arg= dat$experiment, ylim= c(0, max(dat $up)), ylab= 'Internal concentration')

Let's say that if you repeated each experiment many times you would expect most of the results (say 95%) to be within 1 unit away from what you have, then your results would look like: dat <- ame(experiment = c('no glu', 'glu', 'gluFCCP', 'gluV', 'gluN', 'gluNV'), If so, I would present the data as it is adding an acceptable measure of uncertainty. Also, you may have an expectation and explanation for the differences between conditions. Perhaps is accepted in the field that glucose concentration is measured quite reliably and your experimental set up is fairly reproducible. Calculates type-II or type-III analysis-of-variance tables for model objects produced by lm, glm, multinom (in the nnet package), polr (in the MASS package), coxph (in the survival package), coxme (in the coxme pckage), svyglm (in the survey package), rlm (in the MASS package), lmer in the lme4 package, lme in the nlme package. However, you could still have a reasonable guess for how much variation there is around each experimental condition. Anova: Anova Tables for Various Statistical Models Description. By default, R uses traditional dummy coding (also called treatment coding), which works great for regression-style output but can produce weird sums of squares estimates for ANOVA style output. You say each experimental condition is done only once so, as others have said, you cannot estimate the variation around each condition and you cannot derive any p-value. One important consideration when running ANOVAs in R is the coding of factors (in this case, wool and tension).
#Rstudio anova how to#
How to change the code/the test used? I just have one value for the concentration for each experiment. In general, the number of observations should exceed the number of parameters (i.e., groups). Running a model using aov() or lm() does not change anything. The number of groups equals the number of observations, which is more variables than you can afford. The foregoing summary output is a little more explicit. Multiple R-squared: 1, Adjusted R-squared: NaNį-statistic: NaN on 5 and 0 DF, p-value: NA Residual standard error: NaN on 0 degrees of freedom
